source(paste0(getwd(), "/__scripts/helpers.R"))2 Download ESM output from ESGF
2.1 Downloading CMIP6 Earth System Models
The Earth System Grid Federation (ESGF) manages the database for handling climate science data, and supports the Coupled Model Intercomparison Project, which is currently in its sixth iteration (CMIP6).
Today, we will be downloading our model outputs via the ESGF website. We will download the simulations for two models, with the following components:
| Component Name | Variable Name | Description |
|---|---|---|
| Activity ID | ScenarioMIP |
ScenarioMIP |
| Source ID | ACCESS-CM2, IPSL-CM6A-LR |
Model names |
| Experiment ID | SSP2-4.5, SSP5-8.5 historical |
Climate scenario (historical ranges 1850-2014; SSPs range 2015-2100). |
| Model variant label | r1i1p1f1 |
Model variant indicating the realization, initilization method, physics processes, and forcing datasets used in the model simulations. |
| Frequency | mon |
Monthly time-step |
| Variable | tos |
Sea-surface temperature |
For more information on what ESM components mean, see Table 1 in (Schoeman et al. 2023).
2.2 Download wget shell scripts
2.2.2 Filtering ESM components
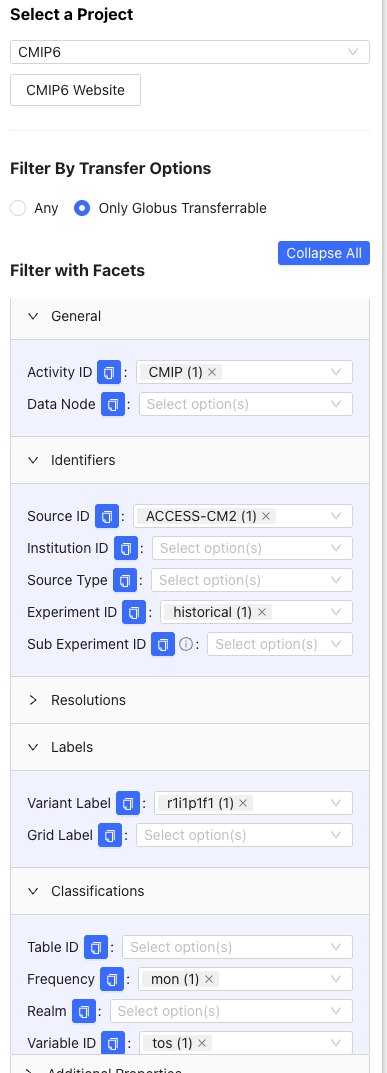
From the left tab, we use the following filters to find the wget script for the ACCESS-CM2 historical simulation:
- Click >General, then under Activity ID, click
CMIP. - Click >Classifications, then under Variable ID, click
tos. Under Frequency, clickmon. - Click >Identifiers, then under Source ID, click
ACCESS-CM2. Under Experiment ID, clickhistorical. - Click >Labels, then under Variant Label, click
r1i1p1f1(this is the most common variant).
2.2.3 Download wget shell script
Then, download the shell script by clicking the Download icon next to the wget download option. This uses wget to download a shell script (i.e., indicated by the .sh file extension) that we will later use to download the file through R whilst interfacing with CDO and the command line.
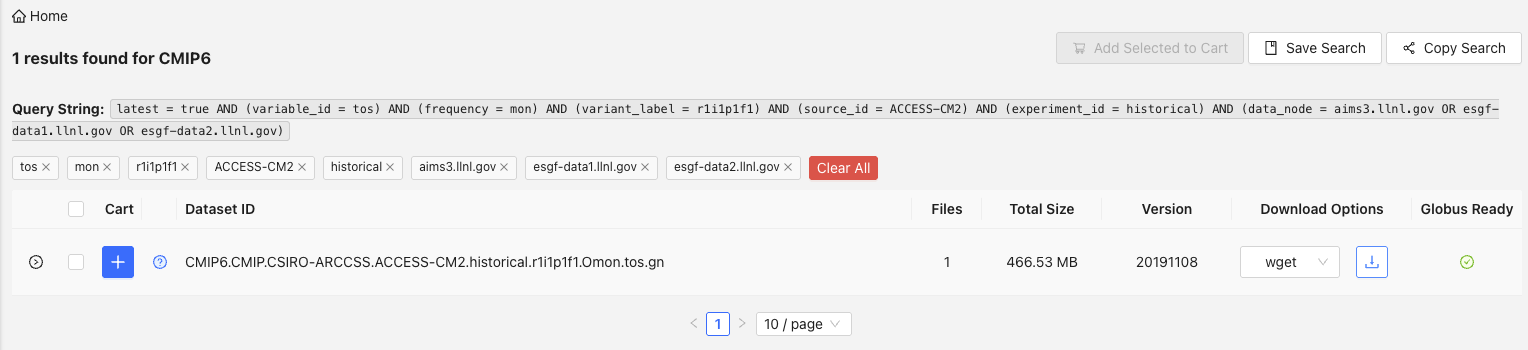
Once downloaded, move the shell script into the /wget_scripts folder in your project directory.
2.2.4 Rinse and repeat
Now, repeat the process for the SSP2-4.5 and SSP5-8.5 scenarios for ACCESS-CM2, in addition to all three scenarios for IPSL-CM6A-LR. You will only need to change the (i) Activity ID from CMIP to ScenarioMIP, and (ii) Experiment ID filter from historical to ssp245 and ssp585, and You will end up with six shell scripts, three for each model.
2.3 Download .nc files with wget scripts
In total, we are about to download eight .nc files that comprise 2.65 GB of data. These are just for two models and two scenarios + historical, at a monthly time-step. Storing ESM outputs can take a lot of space - for example, if you’re working with daily data across multiple models and climate scenarios, you’ll almost certainly need an external hard drive or other storage solution.
We will use wget to download our files from the ESGF Metagrid.
2.3.1 Define dependencies
First, we source our packages and set our directories for where our (i) wget scripts are stored, and (ii) where we will store the ESM outputs.
2.3.2 Prepare for parallel processing
We are harnessing the power of parallel processing to download our ESM outputs. We use availableCores() from the parallelly package to tell us how many available cores we have on our machine. I recommend to use whatever the output is minus two, so that your operating system and other background processes can continue working normally. I have 16 cores on my machine, so I’ll use 14 workers.
parallelly::availableCores() # Output says I have 16 cores on my machine
w <- 14 # Leave two cores for other background processes2.3.3 Function to download files
Now, we write a function that uses the system() function to invoke the terminal from within R to run the wget script, which downloads the ESM file. First, we set our working directory to the full path where data will be stored, which we defined above as /data. Then, we run the system() function to run the wget script using bash via the terminal. Once the script has run and the file has downloaded, we set the working directory back to the root directory.
wget_files <- function(script) {
setwd(paste0(pth, cmip_pth)) #set directory to where data will be stored
system(paste0("bash ", script, " -s")) #run wget on the shell script
setwd(pth) #set directory back to home directory
}Here, we list all wget scripts that we downloaded from ESGF.
files <- list.files(paste0(pth, wget_pth), #full path where wget scripts are stored
pattern = "wget", #only files with wget in the name
full.names = TRUE #list the full path
)
files[1] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-13-19.sh"
[2] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-14-23.sh"
[3] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-16-43.sh"
[4] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-16-6.sh"
[5] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-17-4.sh"
[6] "/Users/admin/Documents/GitHub/esmrworkshop_rladies26/__scripts/wget_scripts/wget_script_2026-2-4_12-5-32.sh" 2.3.4 Run function in parallel
Now, we change to multi-session processing, where multiple workers (sessions) are used to download the files concurrently. We then run the wget_files() function with future_walk() , and use the tictoc package to time how long this takes. Once all files have downloaded, we change back to single-threaded processing.
plan(multisession, workers = w) # Change to multi-threaded processing
tic(); future_walk(files, wget_files); toc() #Run the function in parallel
plan(sequential) # Return to single threaded processing (i.e., sequential/normal)Jessie speed: 30s to 1 min
Download speeds depend on your Wifi connection, the capabilities of your local machine, and the performance of the server/node you’re attempting to connect to. You can check whether ESGF servers are online here.
We can check the files have downloaded by…
list.files(paste0(pth, cmip_pth))[1] "tos_Omon_ACCESS-CM2_historical_r1i1p1f1_gn_185001-201412.nc"
[2] "tos_Omon_ACCESS-CM2_ssp245_r1i1p1f1_gn_201501-210012.nc"
[3] "tos_Omon_ACCESS-CM2_ssp585_r1i1p1f1_gn_201501-210012.nc"
[4] "tos_Omon_IPSL-CM6A-LR_historical_r1i1p1f1_gn_185001-201412.nc"
[5] "tos_Omon_IPSL-CM6A-LR_ssp245_r1i1p1f1_gn_201501-210012.nc"
[6] "tos_Omon_IPSL-CM6A-LR_ssp585_r1i1p1f1_gn_201501-210012.nc" Great! We have successfully downloaded our ESM outputs.
If your scripts have downloaded data for 2100-2300 (i.e., some models will do this), we can remove those files with the rm argument in shell:
system(paste0("rm ", pth, cmip_pth, "/",
"tos_Omon_ACCESS-CM2_ssp585_r1i1p1f1_gn_210101-230012.nc"))
system(paste0("rm ", pth, cmip_pth, "/",
"tos_Omon_IPSL-CM6A-LR_ssp585_r1i1p1f1_gn_210101-230012.nc"))rm
rm is a handy function for removing files, but there is no way of retrieving them if you’ve accidentally removed the wrong file. So, use this function wisely and make sure you’re 100% confident you’re removing the correct file!